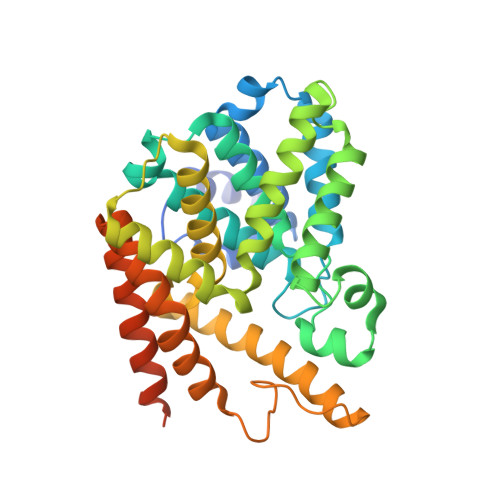
Multiple elements jointly determine inhibitor selectivity of cyclic nucleotide phosphodiesterases 4 and 7
Wang, H., Liu, Y., Chen, Y., Robinson, H., Ke, H.(2005) J Biol Chem 280: 30949-30955
- PubMed: 15994308
- DOI: https://doi.org/10.1074/jbc.M504398200
- Primary Citation of Related Structures:
1ZKL - PubMed Abstract:
Phosphodiesterase (PDE) inhibitors have been widely studied as therapeutics for treatment of human diseases. However, the mechanism by which each PDE family recognizes selectively a category of inhibitors remains a puzzle. Here we report the crystal structure of PDE7A1 catalytic domain in complex with non-selective inhibitor 3-isobutyl-1-methylxanthine and kinetic analysis on the mutants of PDE7A1 and PDE4D2. Our studies suggest at least three elements play critical roles in inhibitor selectivity: 1) the conformation and position of an invariant glutamine, 2) the natures of scaffolding residues, and 3) residues that alter shape and size of the binding pocket. Kinetic analysis shows that single PDE7 to PDE4 mutations increase the sensitivity of PDE7 to PDE4 inhibitors but are not sufficient to render the engineered enzymes comparable with the wild types. The triple S373Y/S377T/I412S mutation of PDE7A1 produces a PDE4-like enzyme, implying that multiple elements must work together to determine inhibitor selectivity.
Organizational Affiliation:
Department of Biochemistry and Biophysics and Lineberger Comprehensive Cancer Center, The University of North Carolina, Chapel Hill, North Carolina 27599-7260, USA.